How to Choose the Right qPCR Probe for Your Experiments?
Choosing the right qPCR probe is crucial for accurate and reliable results in quantitative polymerase chain reaction experiments. According to the market report by Technavio, the global qPCR market is expected to grow by over 10% annually. This increase emphasizes the importance of selecting the most effective qPCR probes to ensure experimental success.
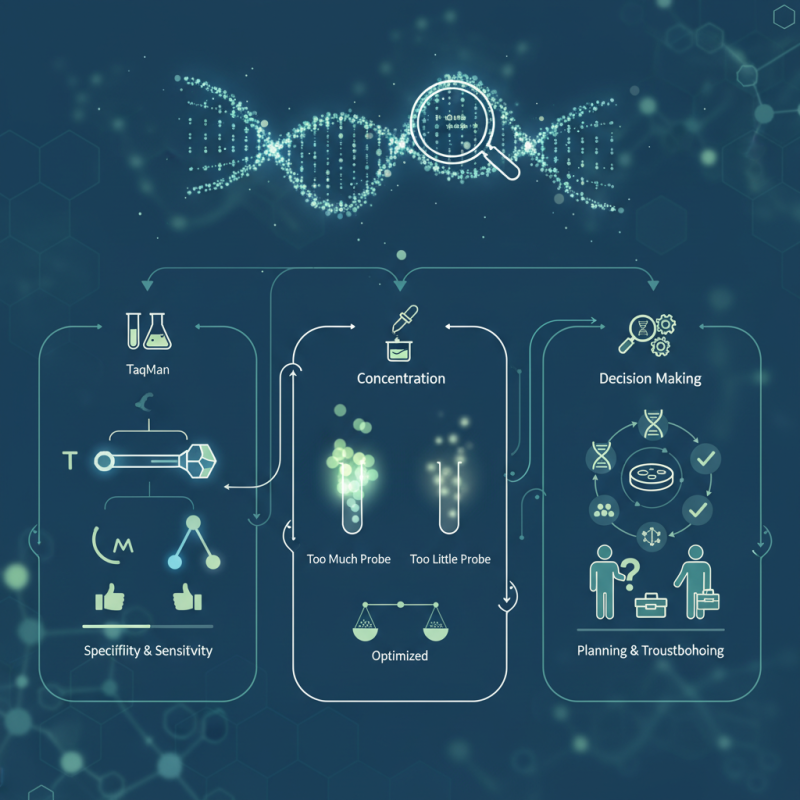
Probes play a vital role in enhancing specificity and sensitivity. A well-chosen qPCR probe minimizes background noise and improves signal detection. Different types of probes, like TaqMan and molecular beacons, have distinct advantages and limitations. Understanding these nuances is essential for researchers seeking robust results.
Moreover, the concentration of the probe can significantly impact the reaction. Excessive probe may lead to signal saturation, whereas too little can result in weak signals. Researchers must consider these variables carefully. Balancing probe design and concentration with the intended application can be challenging. It requires careful planning and troubleshooting. This intricate decision-making process ultimately defines the quality of experimental outcomes.
Understanding the Basics of qPCR Probes
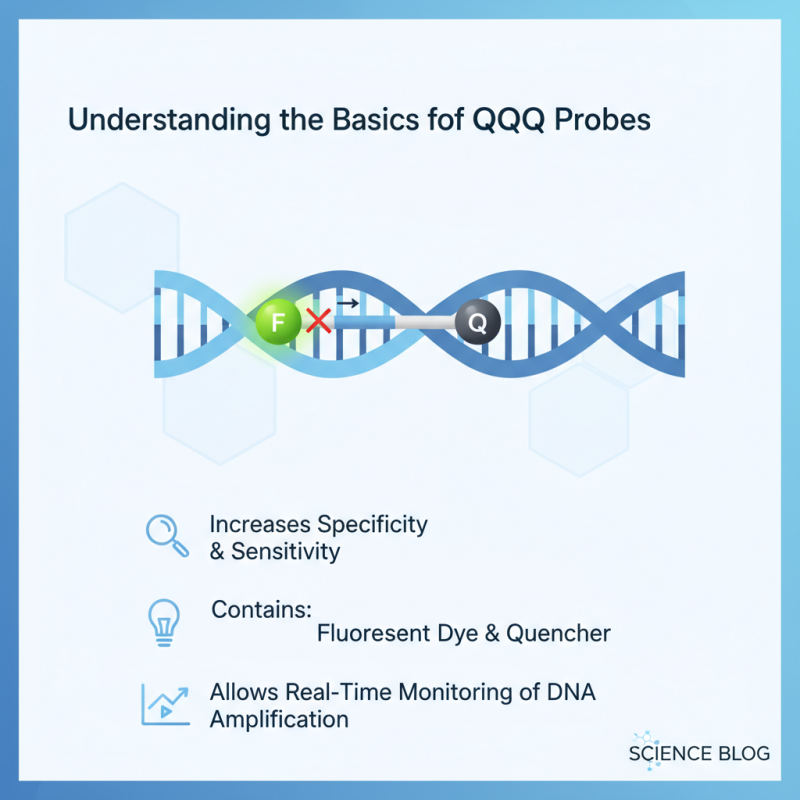

When selecting the right qPCR probe for your experiments, it’s crucial to grasp the fundamentals of qPCR probes. A qPCR probe is used to increase the specificity and sensitivity of your PCR reaction. It typically contains a fluorescent dye and a quencher. This design allows real-time monitoring of DNA amplification.
Consider these tips when choosing your probes. First, ensure your probe's melting temperature (Tm) aligns with your primers. A match of 5-10 degrees Celsius between them is ideal. Second, evaluate the hydrolysis activity of your probe. Some probes may not perform equally, which can impact your results. Be ready to experiment with different options.
Another aspect to reflect on is the importance of specific sequences. High specificity can reduce background noise and improve signal quality. However, even the best probes can sometimes misinterpret the data, especially in complex mixtures. Always validate your results with proper controls to avoid false positives. Trust in your methodology but remain open to adjustments.
Types of qPCR Probes and Their Applications
Choosing the right qPCR probe is crucial for successful experiments. There are several types of qPCR probes, each with unique features. TaqMan probes are popular for their specificity. They work by incorporating a fluorescent dye and a quencher. When the target sequence is amplified, the probe is cleaved, resulting in a detectable fluorescence. These probes are ideal for quantifying gene expression.
Another type is the SYBR Green probe. Unlike TaqMan, SYBR Green binds to double-stranded DNA. It provides a cost-effective option for many researchers. However, it can bind to non-specific products, potentially leading to interpretation challenges. It’s important to validate results, especially when using SYBR Green. Consider using melting curve analysis to enhance specificity.
Lastly, there are molecular beacons. These probes form a hairpin structure that fluoresces upon hybridization with the target. They offer excellent specificity but are more complex to design. Experimenting with different probes can yield varying results. Understanding the strengths and limitations of each type is essential for selecting the best option for your research goals.
Factors to Consider When Selecting a qPCR Probe
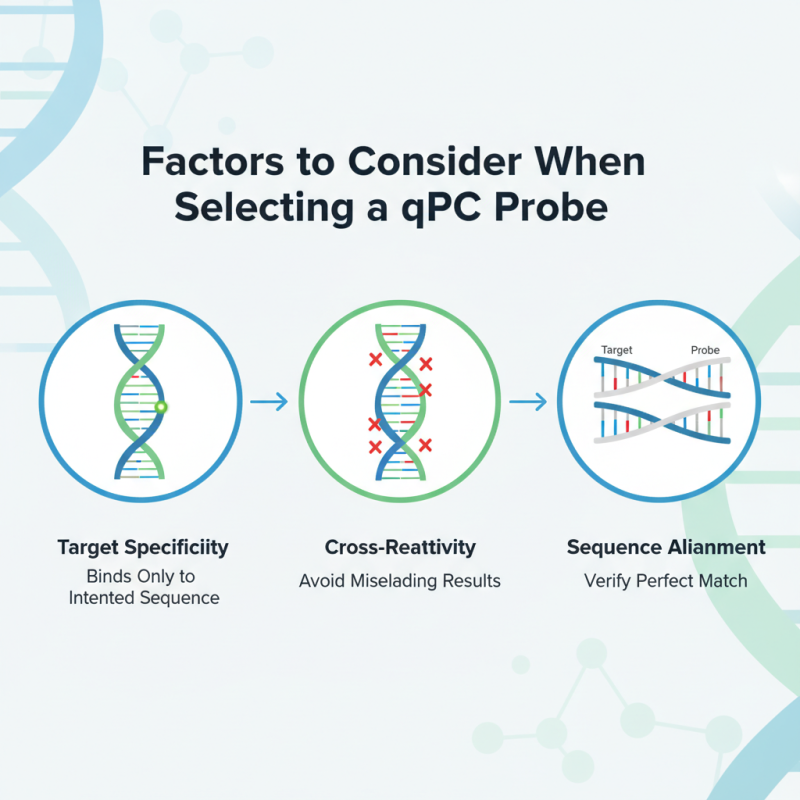
Choosing the right qPCR probe can be daunting. Several factors influence this decision. Target specificity is crucial. A probe should bind only to the intended sequence. Cross-reactivity can lead to misleading results. It’s important to verify that the probe aligns perfectly with your target.
Another key consideration is the probe's melting temperature (Tm). Higher Tm values generally indicate stronger binding. A mismatch can weaken the probe's effectiveness. This may affect your data quality. Think about the assay's sensitivity too. A highly sensitive probe will detect low abundant targets without a hitch.
The design of the probe also deserves attention. Length and GC content impact stability. Generally, a 20-30 nucleotide length works well. Make sure to assess the background signal too. Excessive fluorescence can complicate interpretation. A rigorous evaluation of these parameters can enhance your experiment’s success.
Evaluating Probe Specificity and Sensitivity
Choosing the right qPCR probe requires careful consideration of specificity and sensitivity. Specificity determines how accurately the probe binds to the target sequence. High specificity reduces the chances of cross-reactivity. It's essential to analyze your target sequence and potential off-targets. You may also need to test multiple probes to find the best fit for your experiment.
Sensitivity indicates how well the probe detects low abundance targets. The lower the limit of detection, the more sensitive your probe. Consider the background noise in your samples. A probe with higher sensitivity is beneficial for detecting rare transcripts. However, too high sensitivity might lead to false positives.
Tips for evaluating specificity include designing probes that are unique to the target gene. Use software tools for in-silico analysis. For sensitivity, test different concentrations of the target to find optimum detection limits. Rethink probe binding conditions; they can greatly affect results. Always validate your results with replicates.
Comparison of qPCR Probe Sensitivity and Specificity
This chart illustrates the sensitivity and specificity of different qPCR probes tested in various experiments. The probes are labeled accordingly, and their performance metrics are represented in a bar chart format.
Optimizing Experimental Conditions for qPCR Probes
Optimizing experimental conditions for qPCR probes is crucial for reliable results. Probe design is only one part of the equation. Environmental factors, reagent quality, and amplification protocols also play significant roles. A report from Nature Reviews highlights that inconsistencies in these factors can lead to data variability, affecting the overall accuracy of gene quantification.
To enhance the performance of qPCR assays, consider reaction conditions meticulously. Temperature and cycling times need adjustments based on the specific probe. Optimal annealing temperatures typically range from 55°C to 65°C. Small variations can impact probe binding, affecting assay sensitivity. Additionally, the presence of inhibitors in your samples can skew results. Always ensure you purify RNA or DNA properly before proceeding.
Tips: Regularly validate your probes with controlled experiments. Use positive and negative controls to assess assay performance. It's also beneficial to run a standard curve to quantify your targets accurately. Monitoring these parameters will help you refine techniques, leading to improved reproducibility. However, remember that these optimizations require iterative testing, and it's essential to document all observations meticulously. Balancing precision with practicality is vital.
Related Posts
-

What is rt quantitative pcr and how does it work?
-

2026 Best qPCR Probe Selection Guide for Accurate Results?
-

Top QPCR Probe Types for Accurate Genetic Analysis?
-

How to Choose the Best qPCR Mix for Your Experiments?
-

Why Are qpcr Enzymes Essential for Accurate Gene Expression Analysis?
-
What is PCR Fast and How Does It Work?